Imaging blows up
Cryogenic electron microscopy is driving a burst of discoveries about how molecules work in health and disease

The molecules of life — proteins, DNA and RNA — have many looks.
Some are tubular or spiraled; others are long and spindly, or bulbous and squat. And they’re not stagnant.
Like all molecules, they are a conglomerate of smaller bits — typically, some combination of oxygen, carbon, nitrogen, phosphorous, sulfur and hydrogen atoms connected via bonds of varying strength — and they morph as they perform the many jobs upon which human biology depends.
Even when the body is perfectly still, our insides are moving. That molecular jostle is key to how every cell and organ functions. It’s also a serious challenge for scientists to picture clearly. And so was born a decadeslong quest to capture crisp images of molecules in action.
Early in the field of molecular imaging, the best depictions of molecules were akin to a blurry Rorschach test, showing the overall shape of a molecule in its crystallized state, but not much else.
Now, an imaging technique known as cryogenic electron microscopy, or cryo-EM, helps researchers decipher molecules’ structures in fine detail, even as they flex, twist and undulate.
In a nutshell, the cryo-EM process comes down to roughly four steps: Freeze, shoot, detect, reconstruct. Scientists freeze a molecule of interest, pass a beam of electrons through it, and record how the electrons bounce off the molecule.
That produces images of the molecule, each viewed from a different direction and each molecule caught, frozen, in a certain shape. Researchers can also combine multiple snapshots of a molecule in different conformations, creating movies of how molecules wiggle about, often in different functional states.
Cryo-EM holds several advantages over other molecular imaging methods: It enables investigation of a wider variety of molecules; it allows scientists to see how these molecules exist and act in their natural state or when battling a virus; and the process is relatively fast — if all goes according to plan, a scientist can freeze, shoot and create a model of a molecule in a few days. All of this adds up to a boon for designing drugs and understanding biology and disease.

Cryo-EM has been around since the 1980s, though it wasn’t always the master filmmaker scientists know and love today. With developments in microscope technology and enhanced imaging resolution, it started to gain a little traction in the early 2000s, though by 2002, only eight entries were published in the Electron Microscopy Data Bank, a repository scientists use to collate data about molecular structures.
The boom in cryo-EM-based structure solving took off over the past seven years, thanks to technological advances that not only enhanced its utility as an imaging tool but also made it easier for scientists to use. In 2021 alone, 4,483 cryo-EM structures were deposited in the bank. More than 100 cryo-EM laboratories are spread across the country, and dozens more throughout the world. That may sound like plenty to go around, but for structural biologists, having an electron microscope at your institution is a big deal.
Today, cryo-EM is not just one of the tools for discovering a molecule’s structure — it’s the top tool. Three researchers who developed the technology won the 2017 Nobel Prize in Chemistry for advancing the imaging method.
“Cryo-EM is the most sophisticated technique for studying the structure of proteins,” said Lloyd Minor, MD, dean of the Stanford University School of Medicine. “We made deep investments a number of years ago and essentially went from having almost no presence in cryo-electron microscopy to being an institution that leads in cryo-EM-based research.”
Stanford Medicine obtained five of the multimillion-dollar microscopes operated under the Stanford Cryo-EM Center, and all Stanford faculty have access to the equipment. With these instruments in place, over the past few years the university recruited a number of researchers who excel at this type of structural biology.
The Stanford-associated SLAC National Accelerator Laboratory has an additional five cryo-EM microscopes, with two more on the way, that serve researchers from all over the country.

And it’s not just structural biologists who are invited to use the scopes; cryo-EM experts work with researchers both inside and outside of Stanford to help guide the use of the technology and to help them conduct projects that could benefit from a look at molecular structure.
With the influx of this machinery has come a flurry of new findings by Stanford researchers: Some paint a picture of newly identified lock-and-key binding sites that could be used to guide drug development; some fill in details about the nature of the virus behind COVID-19; and all provide insights about the molecules that make up our bodies and about the diseases that affect them.
“Understanding a protein’s structure allows you to understand its mechanism of action; anything that affects the structure can lead to dysfunction and potentially disease,” said Georgios Skiniotis, PhD, professor of molecular and cellular physiology and of structural biology at Stanford Medicine, professor of photon science at SLAC and scientific director of the Stanford Cryo-EM Center.
Before cryo, there were crystals
Cryo-EM has taken decades to reach its potential. “We used to call cryo-EM ‘blobology’ back in the early 2000s,” said Christopher Barnes, PhD, assistant professor of biology, whose research interests include HIV and SARS-CoV-2, the virus that causes COVID-19.
In its early days, cryo-EM could help scientists understand a protein’s general shape and likeness, but it couldn’t show the details necessary for designing new drugs or fully understanding how the protein interacted with other molecules.
At the time, structural biologists’ preferred imaging method for big biological molecules was X-ray crystallography, the same technique that Rosalind Franklin — a pioneer of structural biology — used to capture the first images of DNA’s twisted-ladder likeness.
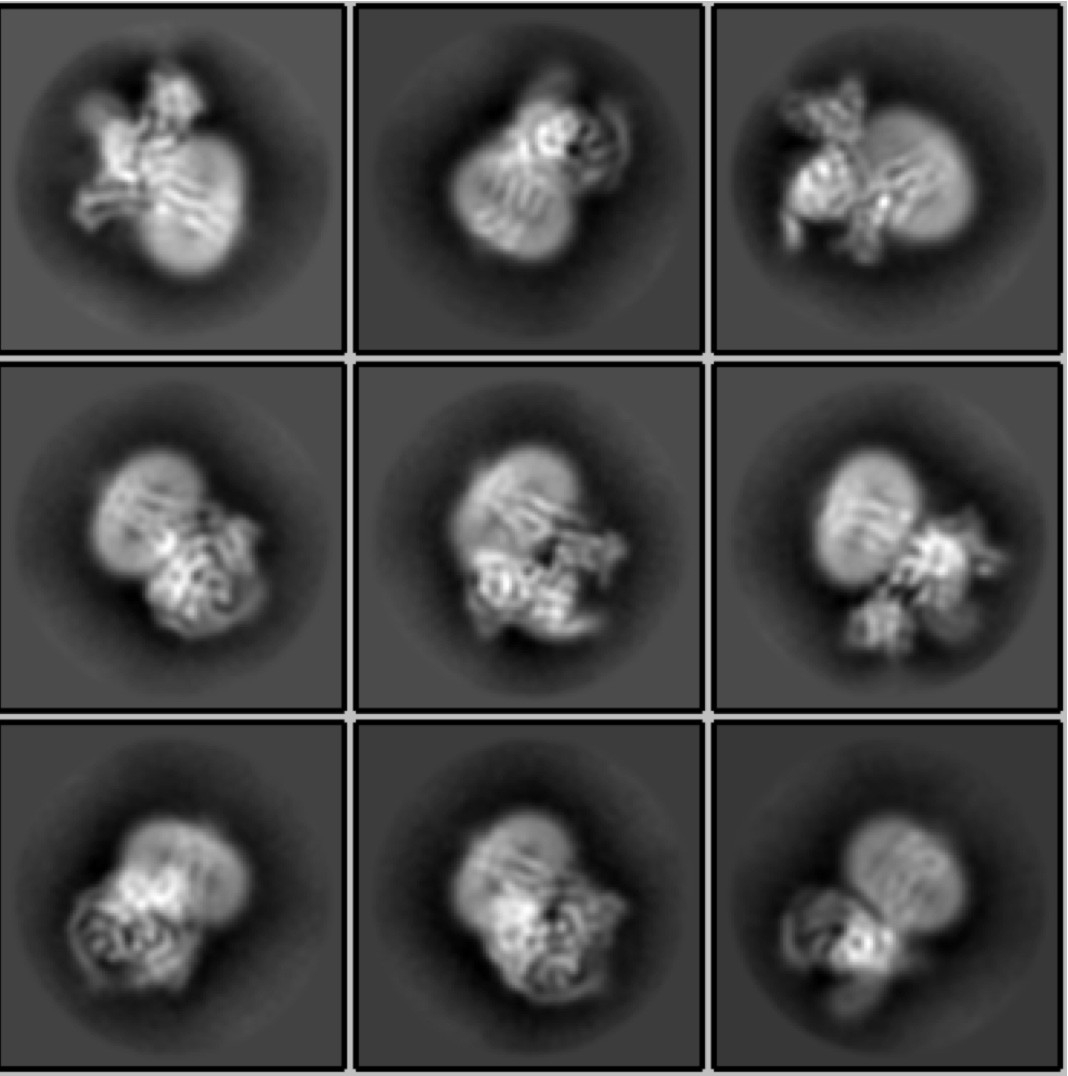
To carry out X-ray crystallography, scientists combine a supersaturated mix of a protein of interest with solutions that help the protein molecules coalesce and eventually form a crystal structure. X-ray beams are shot at the crystal, causing them to diffract, creating an interference pattern that scientists use to infer the structure of the protein.
X-ray crystallography has pitfalls, though, namely, that large molecules are reluctant to form the crystals that are crucial for imaging. Without crystals, there can be no light diffraction, no resulting pattern and no 3D image.
Even if scientists could coax a protein into crystallization, there’s no way to capture its dynamic nature. Instead, it’s caught in a single state — like a tiny Han Solo frozen in its own version of carbonite.
“It only gives you a snapshot of one state of the molecule,” said Skiniotis.
In cryo-EM, millions of proteins swimming in a solution are dropped onto what scientists call a grid — a dish about half the size of a pencil-top eraser composed of thousands of little squares. Liquid ethane, which sits at a crisp minus 165 degrees Celsius or thereabouts, flash freezes the proteins in various conformations.
The scientist then slips the icy tray under the microscope, which shoots it with a beam of electrons (without damaging the molecules), not unlike how scientists shoot X-rays at crystallized proteins during X-ray crystallography.
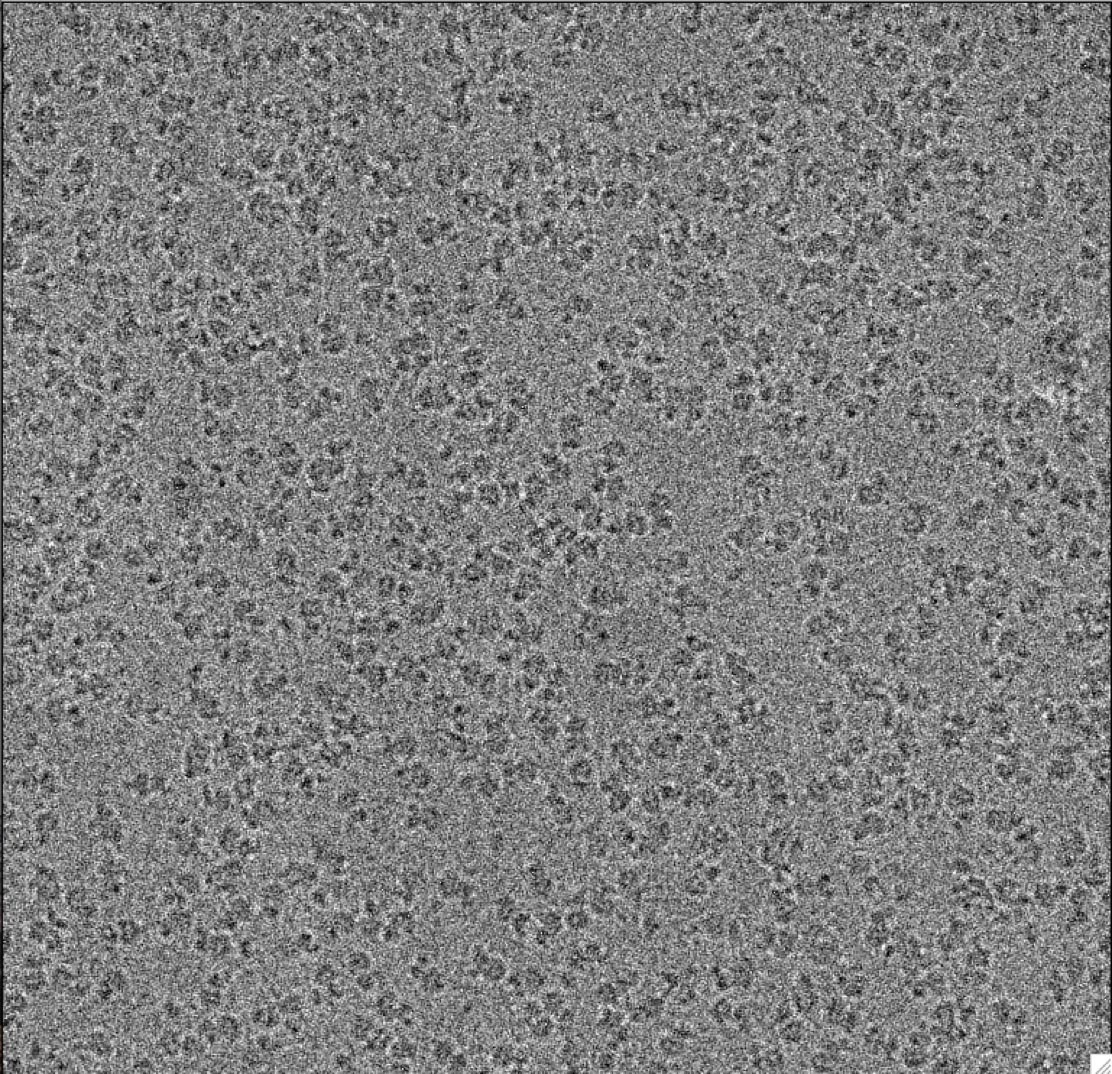
The electrons hit the frozen specimen and ricochet off, creating a pattern of scattered electrons, which an electron detector captures and turns into an image. Each image is a density map that reflects where the highest concentration of molecular material is located (much in the same way a population density map reflects where the most people live).
A few thousand images from different areas of the grid are collected over several hours, each image typically containing two-dimensional maps, called projections, of tens to hundreds of copies of the molecule of interest.

Scientists then use computer software to process the data and reconstruct the three-dimensional shape and structure of the protein. The images are snapshots of copies of a protein, which may be in different conformations, allowing researchers to create movies of protein motion in 3D.
Cryo-EM went through something of a revolution in 2013, when a new era in precise electron detectors, along with leaps in computational power, brought finesse and efficiency to the technique. Armed with new images that reveal a trove of structural information, the science of molecular biology entered its own revolution.
Just as automatic transmissions made cars more accessible to new drivers, advances in detectors and data analysis have expanded access to cryo-EM for structural biologists — and other scientists too.
“You don’t need to know the physics of a microwave to use a microwave,” said Wah Chiu, PhD, a professor of microbiology and immunology and of bioengineering and the director of the Cryo-EM and Bioimaging Division at SLAC. “That’s what it’s like for cryo-EM now. Suddenly, people are solving structures right and left.”
Stanford’s leading cryo-EM researchers are using the technique to develop viral vaccines, devise new viral antidotes and even concoct new treatments for physical and psychic pain.
Know thy enemy
The quest for an HIV vaccine has been underway for decades; but despite intensive research and clinical trials, no HIV vaccine has been approved in the United States. Though treatment options for HIV infection have improved over the past few decades and fewer people die of it, the medicines still have side effects, can be cost-prohibitive, and access is still limited in some countries.
That’s not to say there hasn’t been progress. Recently, researchers in academia and industry have launched trials exploring an mRNA-based vaccine for HIV.
Barnes, a structural biologist recruited for his expertise in cryo-EM, is focusing his research on the envelope glycoprotein, a protein complex that decorates the surface of the virus and plays a crucial role in HIV’s initial infection of cells.
The immune system recognizes that protein complex as an unwanted invader. When the body is infected with HIV, antibodies latch onto the complex and send a signal to the immune system: This molecule is foreign and dangerous; attack. Barnes is using cryo-EM to identify the structures of HIV’s pernicious envelope glycoprotein and how antibodies bind to them.
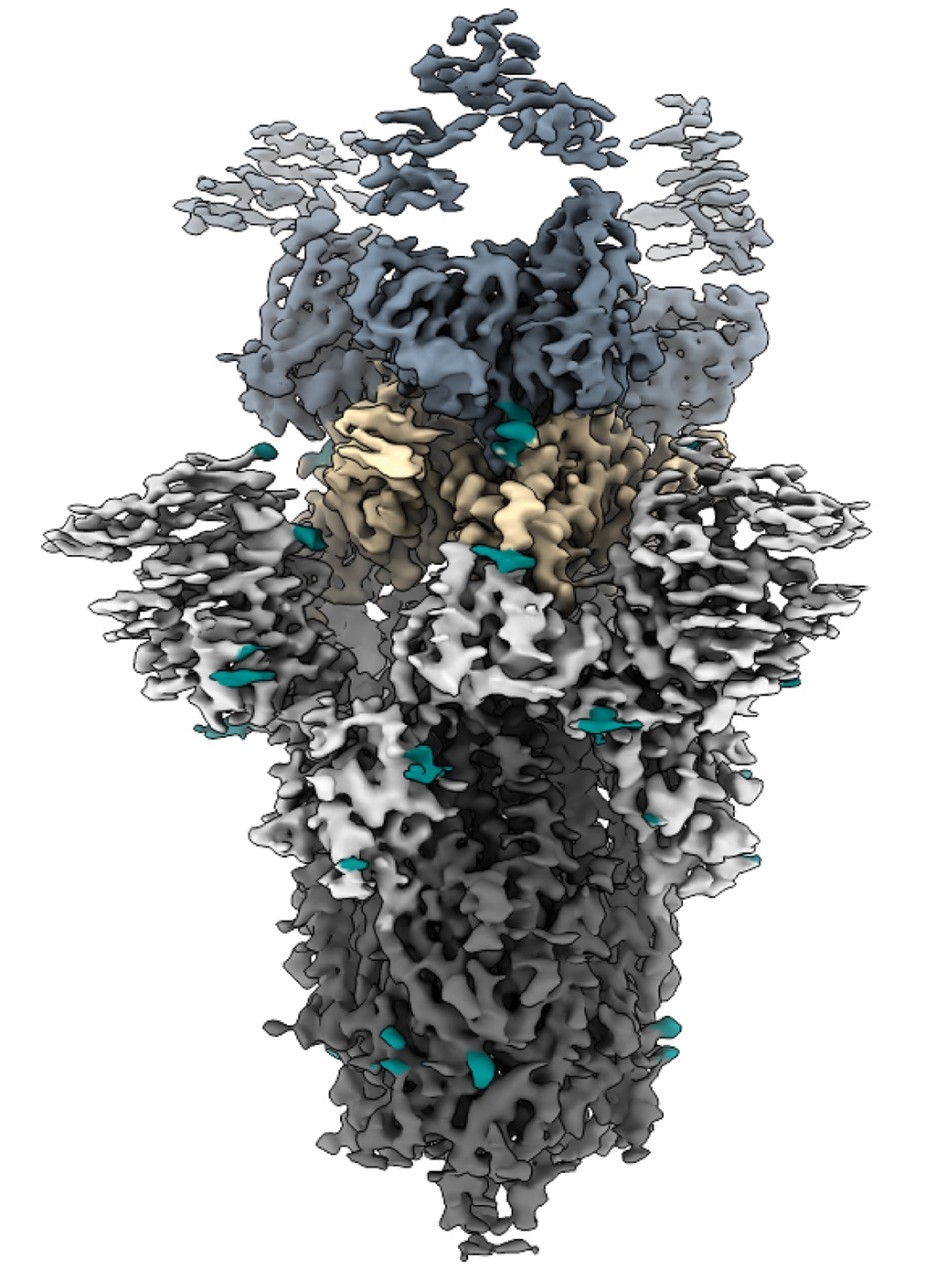
“That’s really important for helping us understand the most potent mechanism to inhibit viral infection,” said Barnes. In particular, he’s interested in one key piece of the protein complex: the region where antibodies bind, known as the epitope.
But there’s more than one type of epitope onto which the antibodies latch, and some epitopes are more conserved than others among the nine existing subtypes of HIV.
“Structural biology can help us identify and characterize those highly conserved regions, which we want to use to design vaccines,” said Barnes. “At the end of the day, you want a vaccine that’s going to induce antibodies that target the most conserved binding regions across HIV strains.”
In past work, Barnes identified antibodies that target an envelope region that’s shared across 70% of HIV varieties, dubbed the silent face — a region that sits across from where the virus attaches itself to human cells in the early stages of infection.
“Now we want to use the structural information we have detailing that region to engineer mimics of the envelope protein to basically elicit a response from the human immune system, such as the generation of broadly neutralizing antibodies, in the hopes of stopping a real infection,” said Barnes.
He hopes that capturing the nooks and crannies of the epitopes in more detail than ever before will finally yield a vaccine that can snuff out all strains of HIV.
Barnes is also zeroing in on coronaviruses, including SARS-CoV-2 and other animal-borne coronaviruses that have the potential to spill over into humans. “We’re trying to identify antibodies that recognize entire families of coronavirus that can have broad disease-fighting capabilities,” he said.
Early in the pandemic, when many people infected with SARS-CoV-2 were donating convalescent plasma for research purposes, Barnes obtained samples to isolate antibodies triggered by the virus. The goal was to use the structural properties of these protective antibodies to inform how to create possible treatments.
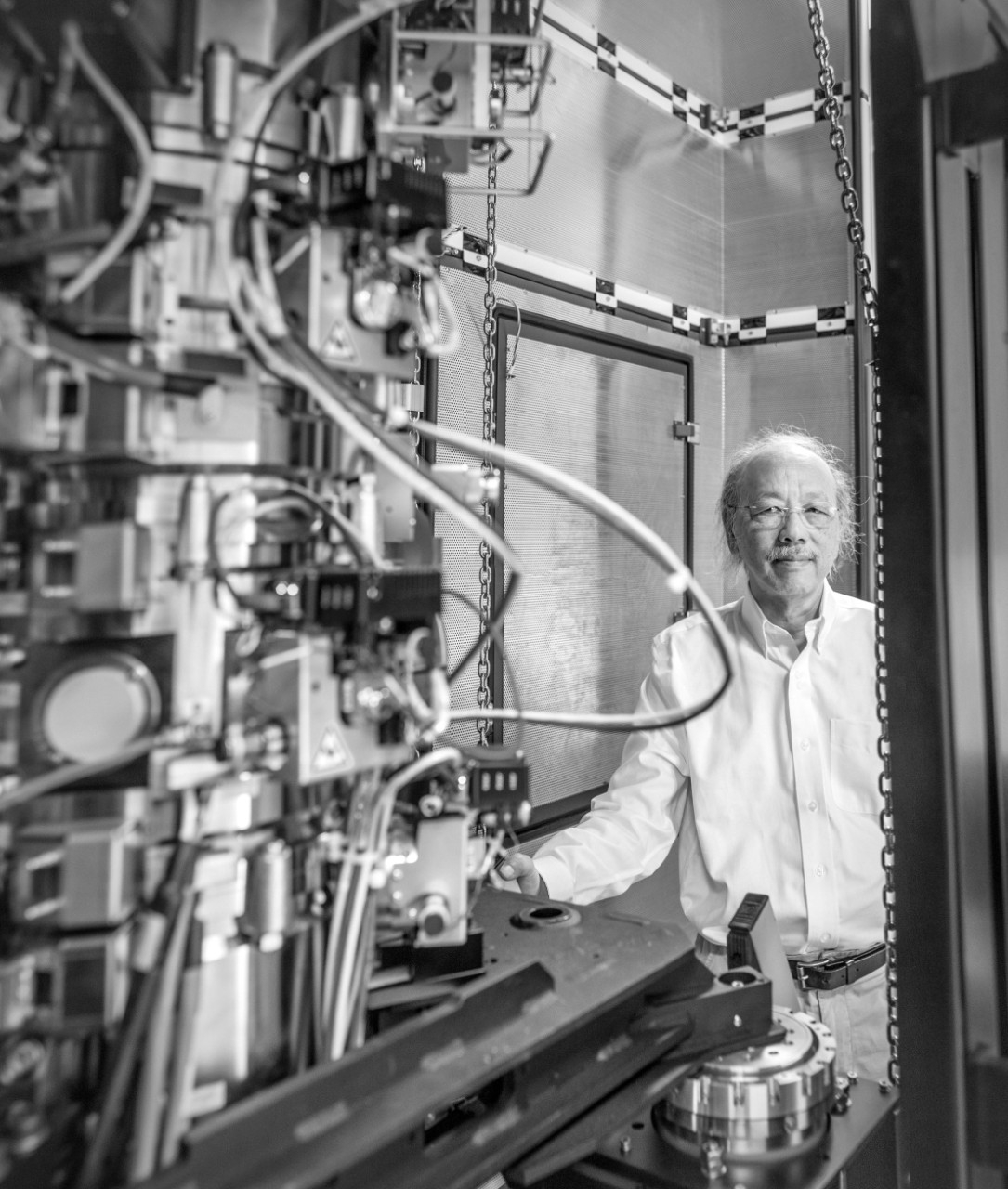

He and colleagues at Rockefeller University identified three neutralizing antibodies with the potential to wipe out COVID-19 infection, two of which are being tested as a treatment. “It was really rewarding to see this work translate to clinical studies that could help to improve outcomes for patients,” Barnes said.
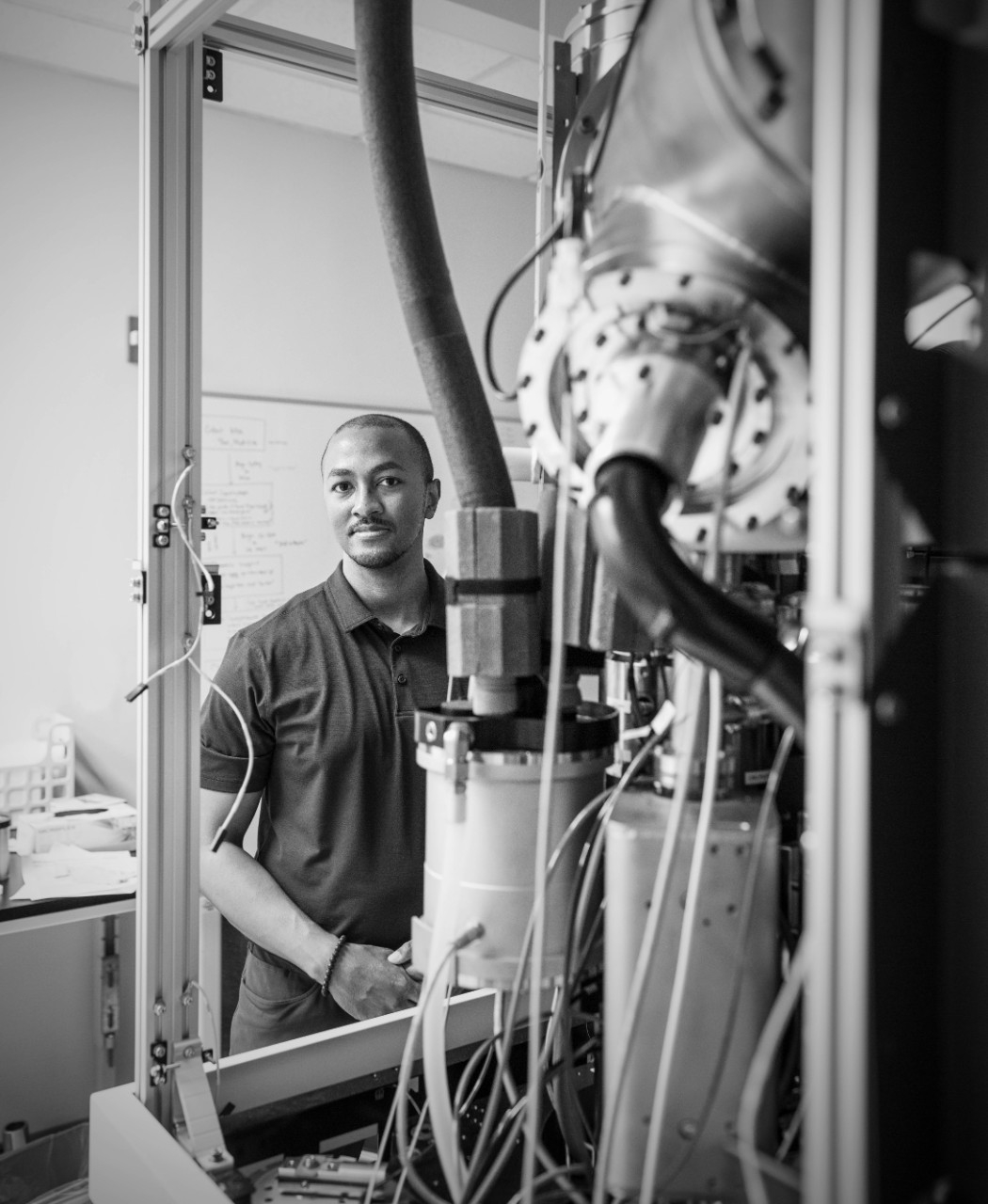
Chiu has also applied his structural expertise to coronaviruses, specifically the infamous spike protein — a name that conjures images of a ball decorated with sturdy spikes that pierce or jab tender cells as it infects.
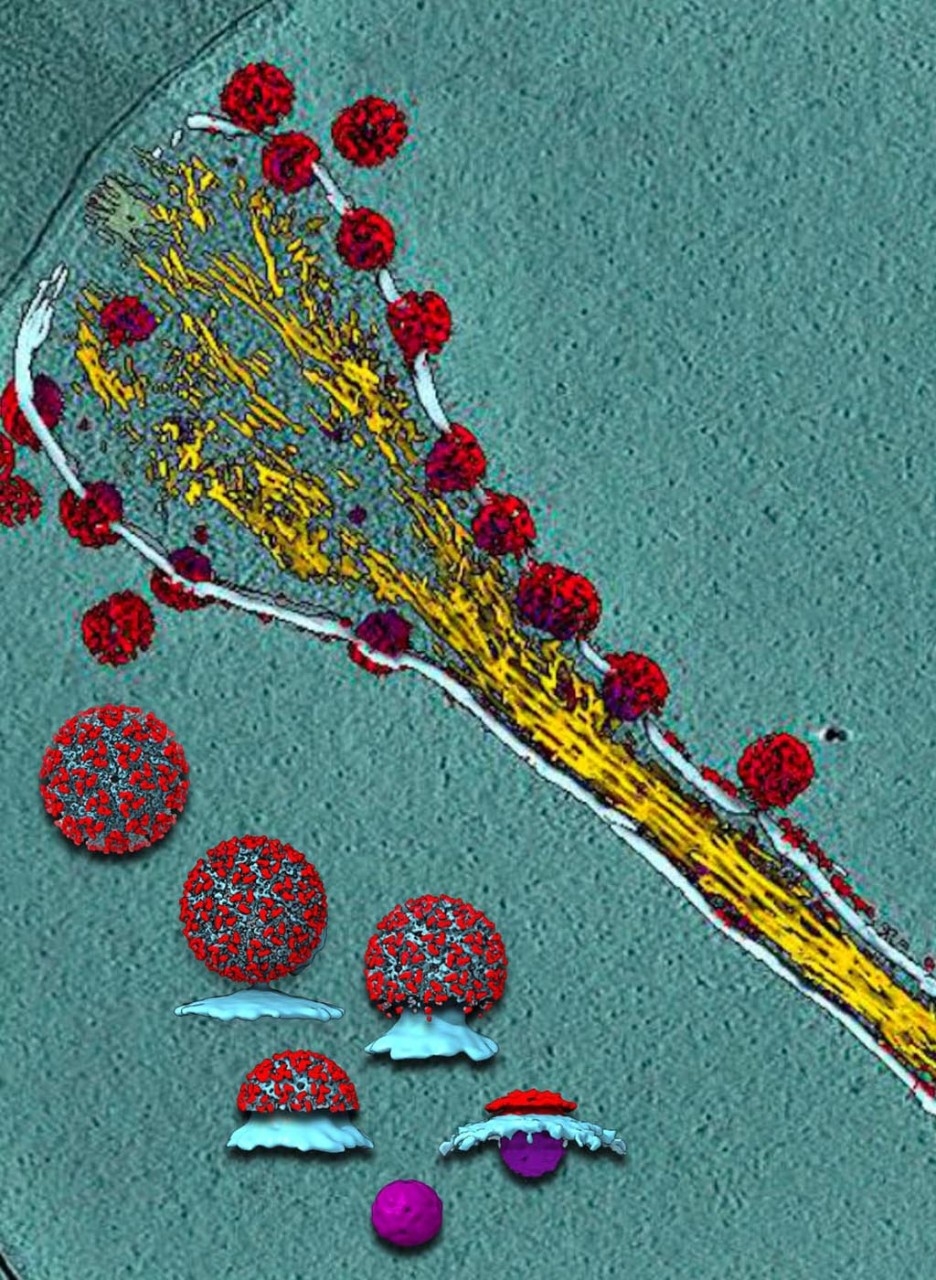
Though they look sharp, cryo-EM imagery has shown what look like spikes aren’t that at all. Instead, they bend, twist, shimmy, shake and jiggle for an unknown purpose.
It’s a mystery how mobility aids the virus’s ability to infect, but the more scientists know about this key protein, especially the unexpected bits (like the bending of the spikes), the better they can understand — and hopefully stop — coronaviruses overall.
Chiu, a pioneer of cryo-EM who’s approaching 50 years of experience using and improving structural biology imaging technology, is also taking a hard look at the mosquito-borne virus that causes chikungunya by zooming out to encompass a whole system of proteins and how they interact with healthy cells to infect humans.
“We’re able to use cryo-EM to create images of a cell that’s mid-infection with chikungunya,” said Chiu. Instead of looking at a single protein of interest, Chiu’s team is harnessing the movie-magic of cryo-EM to watch the chikungunya virus mature and develop inside a cell, and then burst out.
It’s still early days in Chiu’s exploration, but learning more about the structural differences during infection and replication — of key proteins or the virus overall — can shed light on how a virus grows and overtakes a healthy cell.
He and a research team have drawn on structural biology clues to better elucidate how antibodies stop the chikungunya virus from infecting other cells: Antibodies bind to the virus and trigger a cascade of events that stop the virus from multiplying inside the cell, preventing it from exiting the cell to infect others.
The structure of drug design
From behind round-framed glasses, Skiniotis exudes a soft-spoken, animated passion for cryo-EM, his extensive knowledge particularly on display when he talks about his investigation into G-protein-coupled receptors, a family of some of the most diverse and dynamic proteins in the human body.
His drug research centers on these receptors, a class of proteins known for their role in relaying critical signals between cells and tissues to regulate myriad processes, from our heartbeat and mood to our ability to see in a dimly lit room.
These receptors, located on the surface of cells, are known to be the most “druggable” class of receptors — more than 30% of drugs approved by the Food and Drug Administration target these proteins.
But without an understanding of how they contort and connect with neighboring molecules to do their jobs, their drug target potential cannot be fully realized.
“One might take a snapshot of certain proteins coming together and say, ‘I want to break that interaction because it leads to a certain disease. Therefore, I’m going to design a molecule that blocks a central binding site,’” said Skiniotis.
But perhaps blocking that site also affects the binding of other proteins that are beneficial to health, or perhaps it turns out that the drug you designed binds to other off-target proteins. In drug design, the more specific atomic-level information and the more conformations cryo-EM images reveal about a molecule’s structure, the better. That’s what improves drug targeting capabilities, he said.
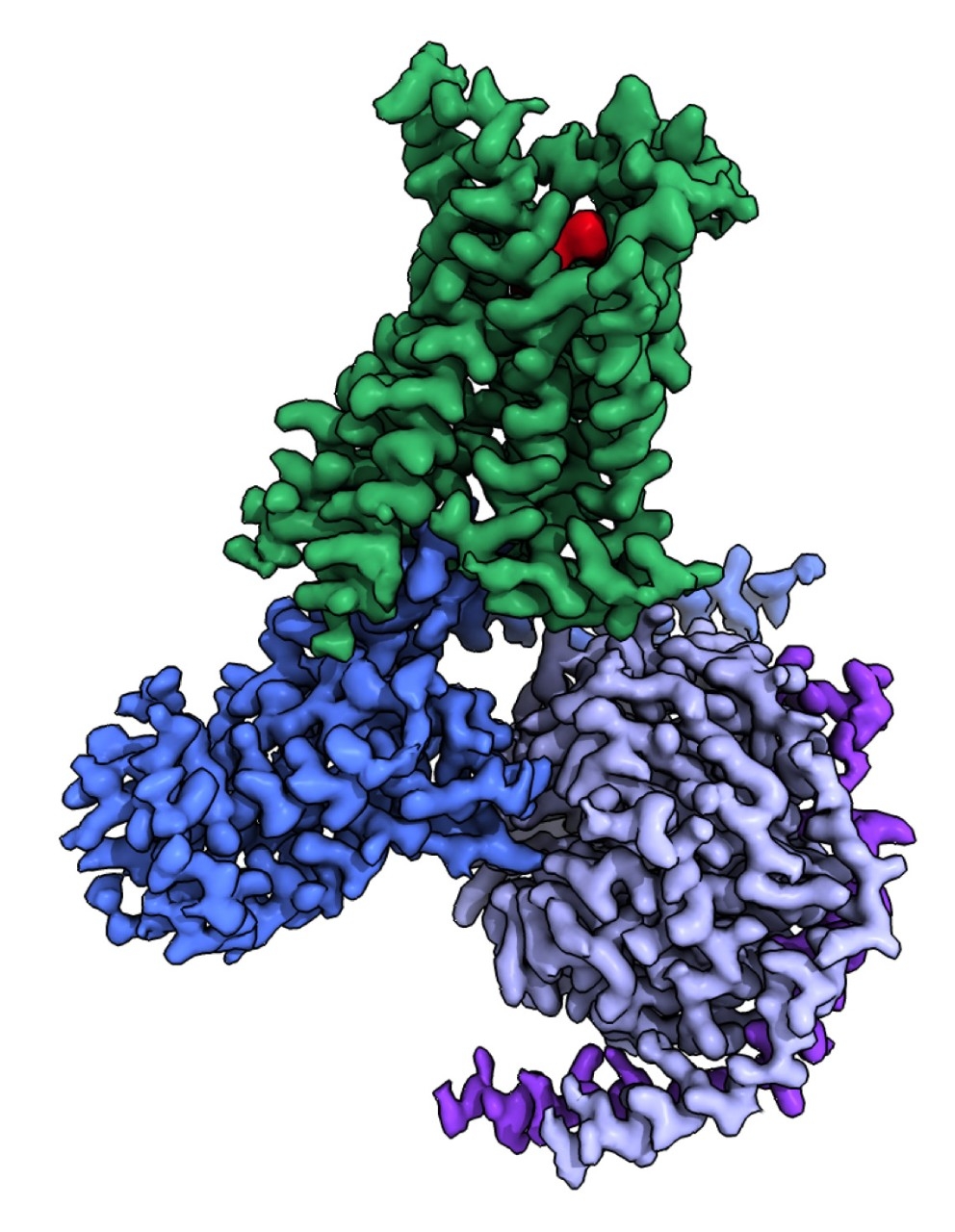
“We have focused on this area for the past few years,” he said. “In my book, you don’t want to get into drug design if you aren’t first focused on mechanisms — drug design requires an understanding of how proteins work and how this function can be blocked, enhanced or modified by certain compounds.”
This is the guiding principle in Skiniotis’ lab as he and his team pursue the next generation of molecules that bind to receptors that trip off a desired cellular response.
“Understanding a protein’s structure allows you to understand its mechanism of action; anything that affects the structure can lead to dysfunction and potentially disease.”
They’re involved in designing these receptor-binding molecules, known as ligands, for three receptor proteins: the opioid receptor, the serotonin receptor and the cannabinoid receptor. Why? Each one has huge potential in treating or assuaging symptoms of human disease, but each also has drawbacks.
“Opioids are great analgesics, but they have severe side effects, the most serious of which is suppression of breathing, which is why people die when they overdose,” said Skiniotis.
But what if the opioid receptor could be targeted to trigger only the pain-mitigating effect, while the molecular pathway that compromises breathing remained silent? That’s why structural specificity and understanding mechanisms — down to the atom — is so crucial for drug design.
“We’re trying to make brand new compounds that elicit only desired effects for human health,” Skiniotis said. That same premise applies to his research on the serotonin receptor. Studies have shown that micro doses of the drug lysergic acid diethylamide, or LSD, which targets the serotonin receptor, can effectively treat post-traumatic stress disorder. “So the question is, can we identify ligands that are good at treating post-traumatic stress disorder by hitting the serotonin receptor without triggering hallucinogenic properties?”
Similarly, he asks whether it might be possible to harness the structural nuance of the cannabinoid receptor to selectively activate the pain-dampening properties without the person feeling intoxicated?
“I cannot tell you, ‘Yes, we have new drugs that will work for sure,’ or even that this idea will be a success in the near future,” said Skiniotis. “But I can tell you that we have new promising and highly specific ligands that we’re testing and that we’re using cryo-EM to continue to finesse the structures of these molecules. Through these ligands, we have started exploring the basic properties of biology so that, perhaps, one day, there will be new drugs that can make a difference.”

